A pair of recent studies from the laboratory of Evangelos Kiskinis, PhD, associate professor in the Ken and Ruth Davee Department of Neurology’s Division of Neuromuscular Disease and of Neuroscience and a member of the Les Turner ALS Center at Northwestern Medicine, have uncovered novel cellular mechanisms that are involved in two types of genetic amyotrophic lateral sclerosis, or ALS.
The findings, published in Science Advances and Cell Reports, improve the understanding of ALS and provide support for the future development of targeted therapies.
‘Baiting’ Genetic Mutations
There are two types of ALS: sporadic (non-genetic), which makes up more than 90 percent of all ALS cases, and familial (genetic).
In their paper published in Science Advances, the Kiskinis team studied a type of familial ALS caused by a repeat genetic sequence of the C9ORF72 gene, which is the largest genetic cause of ALS.
 According to Kiskinis, healthy individuals may have 20 repeats of C9ORF72 in their genome, while patients with ALS can have thousands. These repeats are transcribed and translated through non-canonical pathways which produce irregular RNAs and dipeptides (molecules with two amino acids bonded by a peptide) in neurons.
According to Kiskinis, healthy individuals may have 20 repeats of C9ORF72 in their genome, while patients with ALS can have thousands. These repeats are transcribed and translated through non-canonical pathways which produce irregular RNAs and dipeptides (molecules with two amino acids bonded by a peptide) in neurons.
Kiskinis’ team hypothesized that this cascading effect is what leads to gain-of-function toxicity in motor neurons in this type of ALS. This prompted them to investigate the mechanisms that make these repeated dipeptides (R-DPRs) increasingly toxic.
Using computational and experimental techniques, the investigators found that R-DPRs have a strong binding affinity for RNA molecules. Then, they used a technique called crosslinking immunoprecipitation, or CLIP-seq, to isolate specific RNA fragments and found that R-DPRs bind exclusively to ribosomal RNAs. Specifically, the poly-GR dipeptide binds to ribosomal RNA, which impairs ribosomal homeostasis, a process essential for cell differentiation and the overall makeup of a cell.
Using these findings, the investigators then designed an RNA “bait,” which tricks poly-GR into binding with something that appears to be ribosomal RNA —or in this case, the bait. In both in vivo models and in iPSC neurons derived from patients with C9ORF72 mutations, the bait molecule inhibited toxicity.
“We showed using multiple different approaches that the molecule we designed — the ‘bait’ —binds very highly and specifically to this poly-GR protein, and in binding to it prevents it from binding to ribosomal RNA and prevents it from going into the nucleus, which is where it becomes most toxic,” Kiskinis said. “It serves as a proof of principle to the idea that an RNA molecule that has these ‘bait’ capacities can be therapeutic and it also strengthens the hypothesis that a lot of the toxicity from the C9ORF72 mutation is associated with impairing ribosomal biology.”
The investigators are now optimizing the chemistry of the bait molecule and aim to test it in other models of ALS, according to Kiskinis.
“We’re excited by the fact that iPSC-derived neurons from patients that have these mutations survive a lot longer when we give them this bait molecule, which means that it’s doing something good. We just need to figure out if it could be a viable and transformative therapeutic,” Kiskinis said.
Juan Ortega, PhD, a former postdoctoral fellow in the Kiskinis laboratory, was lead author of the study. Co-authors include Andrew Fleming, a student in the Northwestern University Interdepartmental Neuroscience (NUIN) program, and Samuel Stupp, PhD, the Board of Trustees Professor of Materials Science and Engineering, Chemistry, Medicine, and Biomedical Engineering founding director of the Simpson Querrey Institute for BioNanotechnology (SQI) and the Center for Regenerative Nanomedicine.
This work was supported by the National Institutes of Health (NIH) National Institute of Neurological Disorders and Stroke (NINDS) and National Institute of Aging (NIA) grant R01NS104219; NIH/NINDS grant R21NS107761; the Center for Regenerative Nanomedicine at the Simpson Querrey Institute; the Les Turner ALS Foundation; and the and New York Stem Cell Foundation.
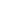
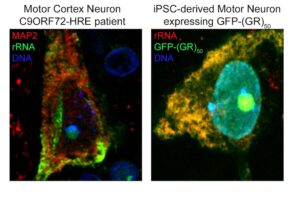 Above left: Neuron at the motor cortex of a C9ORF72 patient stained for MAP2 and rRNA, showing the pattern of expression of ribosomal RNA in a human neuron mostly at the cytosol and the nucleolus. Above right: iPSC-derived motor neuron transduced with GFP-(GR)50 lentivirus, and stained for rRNA, showing the total overlap of poly-GR and rRNA in human motor neurons. Courtesy of Juan Ortega, PhD.
Above left: Neuron at the motor cortex of a C9ORF72 patient stained for MAP2 and rRNA, showing the pattern of expression of ribosomal RNA in a human neuron mostly at the cytosol and the nucleolus. Above right: iPSC-derived motor neuron transduced with GFP-(GR)50 lentivirus, and stained for rRNA, showing the total overlap of poly-GR and rRNA in human motor neurons. Courtesy of Juan Ortega, PhD.
Linking Causal Genes
There are 30 known genes associated with familial forms of ALS, but whether each genetic driver contributes to just one general type of ALS or multiple types has remained a looming question in the field, according to Kiskinis.
To address this question, according to their study published in Cell Reports, Kiskinis’ team used patient-derived induced-pluripotent-stem-cell (iPSC) spinal motor neurons to develop a model of the second most common type of familial ALS, which is caused by mutations in the SOD1 gene and is apparent in two percent of all ALS cases.
“We know from work that’s been done in the field over the last 20 years, that this mutation causes toxicity through gain-of-function effects. We’ve known that mutations lead to protein misfolding, and the misfolding starts a cascade that leads to dysfunction. But how that happens we don’t quite understand,” Kiskinis said.
The investigators used temporal proteomics to compare the proteomes of control iPSC neurons and iPSC neurons with a SOD1 mutation and compared the rate of protein degradation (the recycling and replacing of old proteins) between both sets of neurons.
In the mutant-SOD1 iPSC neurons, they found that another ALS causal protein — valosin-containing protein (VCP) — degraded slower than in the isogenic control iPSC neurons. This slow protein degradation ultimately caused VCP to interact less with certain proteins and more with other proteins.
“This is quite exciting because mutations in VCP that can cause ALS are thought to act in a similar way,” Kiskinis said.
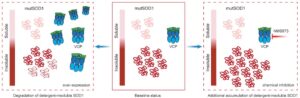 Above: When VCP is overexpressed (left) in the context of mutant SOD1, the levels of detergent-insoluble SOD1 protein are decreased. In contrast, upon chemical inhibition of VCP with NMS873 (right), there is significant accumulation of SOD1 protein within the detergent-insoluble fraction. Courtesy of Konstantinos Tsioras, PhD.
Above: When VCP is overexpressed (left) in the context of mutant SOD1, the levels of detergent-insoluble SOD1 protein are decreased. In contrast, upon chemical inhibition of VCP with NMS873 (right), there is significant accumulation of SOD1 protein within the detergent-insoluble fraction. Courtesy of Konstantinos Tsioras, PhD.
The investigators then aimed to determine whether this altered VCP function played a role in SDO1-mediated toxicity. In both mutant-SOD1 iPSC-patient neurons and C.elegans (a type of roundworm) models of mutant SOD1, the investigators found that when VCP was overexpressed, toxicity decreased and when VCP function was inhibited, toxicity increased.
“This paper, and work we’ve done in our lab over the last few years, are demonstrating that while the genetic causes might be different, there’s always some sort of level of mechanistic overlap when it comes to the causes of dysfunction in these cells,” Kiskinis said.
This is important, Kiskinis added, because the findings can inform the design of future clinical trials and improve the application of targeted therapeutics. Going forward, Kiskinis said his team wants to determine if VCP is an effective therapeutic target for patients with a SOD1 mutation, as well as for other types of ALS.
Konstantinos Tsioras, PhD, a postdoctoral fellow in the Kiskinis laboratory, was the lead author of the study. Coauthors include Robert Kalb, MD, the Joan and Paul Rubschlager Professor and director of the Les Turner ALS Center, and Jeffrey Savas, PhD, assistant professor in the Ken and Ruth Davee Department of Neurology’s Division of Behavioral Neurology.
This work was supported by the National Institutes of Health (NIH), National Institute of Neurological Disorders and Stroke (NINDS), National Institute on Aging R01NS104219, NIH/NINDS grants R21NS107761, the Les Turner ALS Foundation, and the New York Stem Cell Foundation.
Originally published at the Northwestern Feinberg School of Medicine.

